MIT researchers reported in Nature that chromatin moves in two distinct ways across cell types, based on MINFLUX tracking in living cells. The findings may help explain how cells search for gene-regulating partners and repair DNA damage.
MIT researchers have mapped chromatin movement in living cells and identified two distinct motion modes that may help explain how gene expression and DNA repair are controlled.
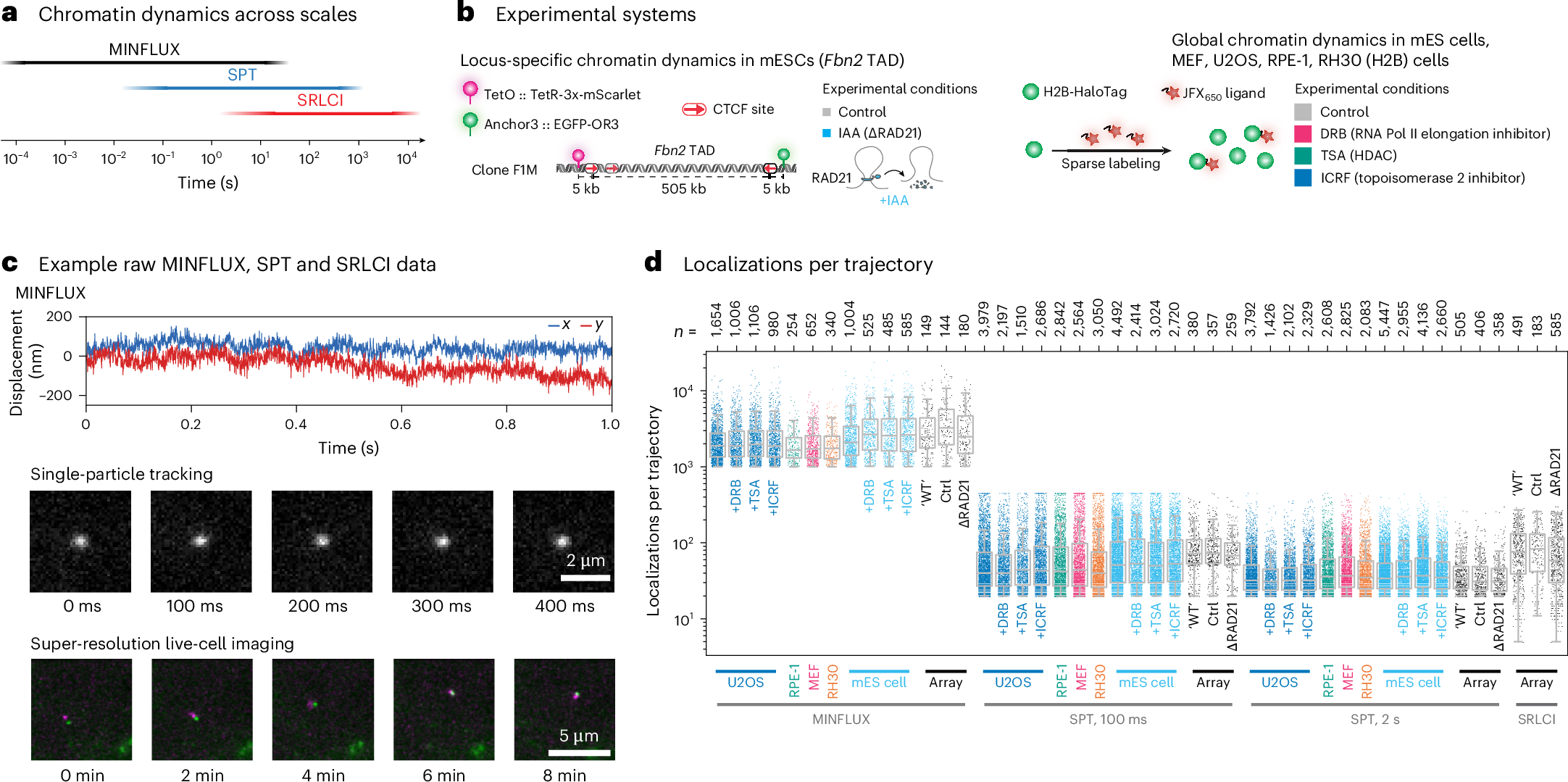
The study, published in Nature on May 4, 2026, used MINFLUX-based tracking to follow chromatin across five mouse and human cell types over a time span covering seven orders of magnitude. MIT News said the work shows chromatin does not move in a single uniform way, but instead falls into two dynamics classes.
According to the researchers, those motion patterns matter because chromatin movement influences how the cell searches for enhancer-promoter contacts and how it responds to DNA double-strand breaks. That makes the work relevant not just to basic chromosome biology, but also to processes tied to gene regulation and genome maintenance.
MIT’s companion summary said the findings provide a new framework for thinking about how chromatin behavior varies across cell types. Phys.org later republished the same core findings.
The research adds detail to a long-running question in cell biology: how the physical movement of DNA inside the nucleus helps determine when genes are turned on and how damage is repaired.
Revision note
Initial automated publication.